To better understand and perhaps prevent cancers brought on by multiple genetic mutations, Rice University researchers are constructing a theoretical framework.
A new theory suggests that mutations have few straightforward ways to establish themselves in cells and cause tumors.
For many researchers, the road to cancer prevention is long and difficult, but a recent study by Rice University scientists suggests that there may be shortcuts.
A theoretical framework is being developed by Rice scientist Anatoly Kolomeisky, postdoctoral researcher Hamid Teimouri, and research assistant Cade Spaulding that will explain how cancers brought on by several genetic mutations might be more readily recognized and perhaps prevented.
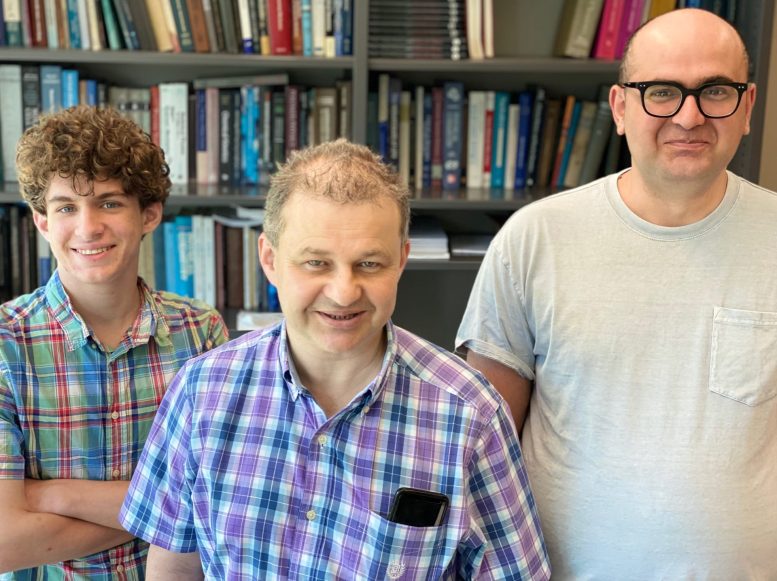

A new paper by a Rice University lab shows how to increase the odds of identifying cancer-causing mutations before tumors take hold. Authors are, from the left, Cade Spaulding, Anatoly Kolomeisky, and Hamid Teimouri. Credit: Rice University
It does this by detecting and ignoring transition pathways that don’t significantly contribute to the fixation of mutations in a cell that later becomes a tumor.
The study, which was published on May 13th, 2022 in the Biophysical Journal, details their analysis of the effective energy landscapes of cellular transformation pathways connected to a number of cancers. The ability to narrow the number of paths to those most likely to initiate cancer could help in the development of strategies to interrupt the process before it begins.
“In some sense, cancer is a bad-luck story,” said Kolomeisky, a professor of chemistry and of chemical and biomolecular engineering. “We think we can decrease the probability of this bad luck by looking for low-probability collections of mutations that typically lead to cancer. Depending on the type of cancer, this can range between two mutations and 10.”
Calculating the effective energies that govern interactions in biomolecular systems may help anticipate how they will behave. The theory is widely used to anticipate how a protein will fold based on the sequence of its constituent atoms and how they interact.
The Rice team is applying the same idea to cancer initiation pathways that work in cells but sometimes include mutations that are undetected by the body’s protections. When two or more of these mutations are fixed in a cell, they are carried on when cells divide and tumors develop.

An algorithm developed at Rice University identifies and ignores transition pathways that don’t contribute much to the fixation of mutations in a cell that goes on to establish a tumor. Credit: Hamid Teimouri/Rice University
By their calculations, the odds favor the most dominant pathways, those that carry mutations forward while expending the least amount of energy, Kolomeisky said.
“Instead of looking at all possible chemical reactions, we identify the few that we might need to look at,” he explained. “It seems to us that most tissues involved in the initiation of cancer are trying to be as homogenous as possible. The rule is a pathway that decreases heterogeneity is always going to be the fastest on the road to tumor formation.”
The huge number of possible pathways seems to make narrowing them down an intractable problem. “But it turned out that using our chemical intuition and building an effective free-energy landscape helped by allowing us to calculate where in the process a mutation is likely to become fixated in a cell,” Kolomeisky said.
The team simplified calculations by focusing initially on pathways involving only two mutations that, when fixed, initiate a tumor. Kolomeisky said mechanisms involving more mutations will complicate calculations, but the procedure remains the same.
Much of the credit goes to Spaulding, who under Teimouri’s direction created the algorithms that greatly simplify the calculations. The visiting research assistant was 12 when he first met Kolomeisky to ask for guidance. Having graduated from a Houston high school two years early, he joined the Rice lab last year at 16 and will attend Trinity University in San Antonio this fall.
“Cade has outstanding ability in computer programming and in implementing sophisticated algorithms despite his very young age,” Kolomeisky said. “He came up with the most efficient Monte Carlo simulations to test our theory, where the size of the system can involve up to a billion cells.”
Spaulding said the project brought together chemistry, physics and biology in a way that meshes with his interests, along with his computer programming skills. “It was good way to combine all of the branches of science and also programming, which is what I find most interesting,” he said.
The study follows a 2019 paper in which the Rice lab modeled stochastic (random) processes to learn why some cancerous cells overcome the body’s defenses and trigger spread of the disease.
But understanding how those cells become cancerous in the first place could help head them off at the pass, Kolomeisky said. “This has implications for personalized medicine,” he said. “If a tissue test can find mutations, our framework might tell you if you are likely to develop a tumor and whether you need to have more frequent checkups. I think this powerful framework can be a tool for prevention.”
The Welch Foundation (C-1559), the National Science Foundation (1953453, 1941106) and the NSF-supported Center for Theoretical Biological Physics (2019745) supported the research.
Reference: “Optimal pathways control fixation of multiple mutations during cancer initiation” by Hamid Teimouri, Cade Spaulding and Anatoly B. Kolomeisky, 13 May 2022, Biophysical Journal.
DOI: 10.1016/j.bpj.2022.05.011

0Comments